Single detector, software injection (linear polarisation)#
Here we compare lalpulsar_parameter_estimation_nested with cwinpy in the case of
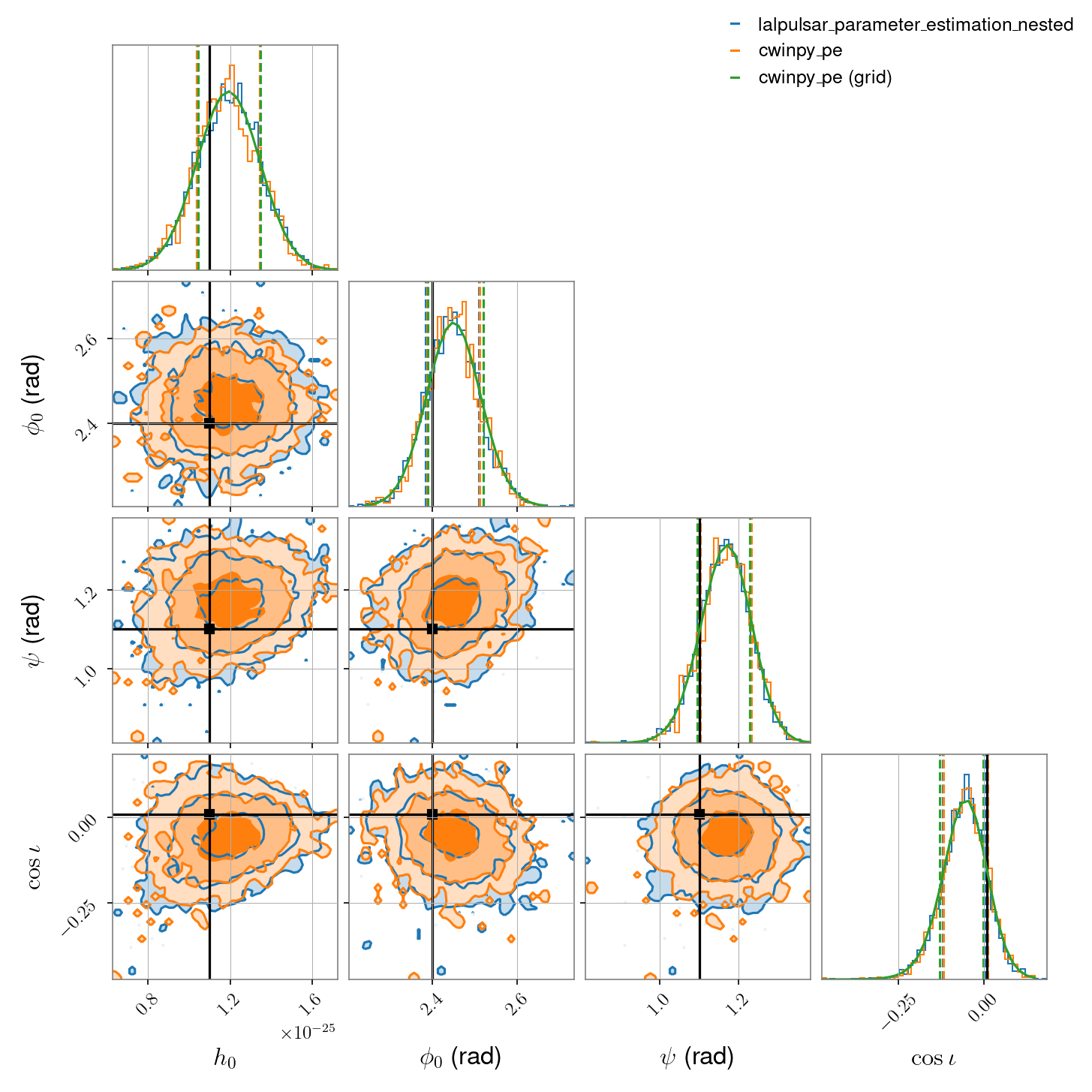
simulated Gaussian noise from a single detector (H1 in this case) containing a software injected signal with close-to linear polarisation. The parameters being
estimated are \(h_0\), \(\phi_0\), \(\psi\) and \(\cos{\iota}\), all with uniform priors.
The script for this comparison, using the dynesty nested sampling algorithm, is shown at the bottom of the page. It produces the following comparison data:
Method |
\(\\ln{(Z)}\) |
\(\\ln{(Z)}\) noise |
\(\\ln{}\) Odds |
|---|---|---|---|
|
162961.652 |
162940.311 |
21.341±0.107 |
|
162961.875 |
162940.311 |
21.564±0.185 |
|
162961.593 |
21.282 |
Method |
\(h_0\) |
\(\\phi_0\) (rad) |
\(\\psi\) (rad) |
\(\\cos{\\iota}\) |
|---|---|---|---|---|
|
1.19±0.16×10-25 |
2.45±0.06 |
1.16±0.06 |
-0.06±0.07 |
90% credible intervals |
[0.94, 1.45]×10-25 |
[2.34, 2.56] |
[1.06, 1.27] |
[-0.17, 0.05] |
|
1.18±0.15×10-25 |
2.45±0.06 |
1.17±0.07 |
-0.06±0.07 |
90% credible intervals |
[0.92, 1.44]×10-25 |
[2.35, 2.56] |
[1.06, 1.28] |
[-0.17, 0.05] |
Method |
\(h_0\) |
\(\\phi_0\) (rad) |
\(\\psi\) (rad) |
\(\\cos{\\iota}\) |
\(\\ln{(L)}\) max |
|---|---|---|---|---|---|
|
1.25×10-25 |
2.44 |
1.17 |
-0.05 |
162975.01 |
|
1.22×10-25 |
2.44 |
1.17 |
-0.05 |
162975.02 |
#!/usr/bin/env python
"""
Compare cwinpy with lalpulsar_parameter_estimation_nested for
data from a single detector detector containing a software injection
with close-to-linear polarisation.
"""
import os
import subprocess as sp
import h5py
import matplotlib
import numpy as np
from bilby.core.prior import Uniform
from comparitors import comparisons
from lalinference import LALInferenceHDF5PosteriorSamplesDatasetName
from lalinference.io import read_samples
from matplotlib import pyplot as plt
from solar_system_ephemerides.paths import body_ephemeris_path, time_ephemeris_path
from cwinpy import HeterodynedData
from cwinpy.pe import pe
from cwinpy.plot import Plot
matplotlib.use("Agg")
# create a fake pulsar parameter file
parcontent = """\
PSRJ J0123+3456
RAJ 01:23:45.6789
DECJ 34:56:54.321
F0 567.89
F1 -1.2e-12
PEPOCH 56789
H0 1.1e-25
COSIOTA 0.01
PSI 1.1
PHI0 2.4
"""
injection_parameters = {}
injection_parameters["h0"] = 1.1e-25
injection_parameters["phi0"] = 2.4
injection_parameters["psi"] = 1.1
injection_parameters["cosiota"] = 0.01
label = "single_detector_software_injection_linear"
outdir = "outputs"
if not os.path.isdir(outdir):
os.makedirs(outdir)
# add content to the par file
parfile = os.path.join(outdir, "{}.par".format(label))
with open(parfile, "w") as fp:
fp.write(parcontent)
# create some fake heterodyned data
detector = "H1" # the detector to use
asd = 1e-24 # noise amplitude spectral density
times = np.linspace(1000000000.0, 1000086340.0, 1440) # times
het = HeterodynedData(
times=times,
par=parfile,
injpar=parfile,
inject=True,
fakeasd=asd,
detector=detector,
)
# output the data
hetfile = os.path.join(outdir, "{}_data.txt".format(label))
het.write(hetfile)
# create priors
phi0range = [0.0, np.pi]
psirange = [0.0, np.pi / 2.0]
cosiotarange = [-1.0, 1.0]
h0range = [0.0, 1e-23]
# set prior for lalpulsar_parameter_estimation_nested
priorfile = os.path.join(outdir, "{}_prior.txt".format(label))
priorcontent = """H0 uniform {} {}
PHI0 uniform {} {}
PSI uniform {} {}
COSIOTA uniform {} {}
"""
with open(priorfile, "w") as fp:
fp.write(priorcontent.format(*(h0range + phi0range + psirange + cosiotarange)))
# set prior for bilby
priors = {}
priors["h0"] = Uniform(h0range[0], h0range[1], "h0", latex_label=r"$h_0$")
priors["phi0"] = Uniform(
phi0range[0], phi0range[1], "phi0", latex_label=r"$\phi_0$", unit="rad"
)
priors["psi"] = Uniform(
psirange[0], psirange[1], "psi", latex_label=r"$\psi$", unit="rad"
)
priors["cosiota"] = Uniform(
cosiotarange[0], cosiotarange[1], "cosiota", latex_label=r"$\cos{\iota}$"
)
# run lalpulsar_parameter_estimation_nested
try:
execpath = os.environ["CONDA_PREFIX"]
except KeyError:
raise KeyError(
"Please work in a conda environment with lalsuite and cwinpy installed"
)
execpath = os.path.join(execpath, "bin")
lppen = os.path.join(execpath, "lalpulsar_parameter_estimation_nested")
n2p = os.path.join(execpath, "lalinference_nest2pos")
Nlive = 1000 # number of nested sampling live points
Nmcmcinitial = 0 # set to 0 so that prior samples are not resampled
outfile = os.path.join(outdir, "{}_nest.hdf".format(label))
# set ephemeris files
efile = body_ephemeris_path(body="earth", jplde="DE405")
sfile = body_ephemeris_path(body="sun", jplde="DE405")
tfile = time_ephemeris_path(units="TCB")
# set the command line arguments
runcmd = " ".join(
[
lppen,
"--verbose",
"--input-files",
hetfile,
"--detectors",
detector,
"--par-file",
parfile,
"--prior-file",
priorfile,
"--Nlive",
"{}".format(Nlive),
"--Nmcmcinitial",
"{}".format(Nmcmcinitial),
"--outfile",
outfile,
"--ephem-earth",
str(efile),
"--ephem-sun",
str(sfile),
"--ephem-timecorr",
str(tfile),
]
)
with sp.Popen(
runcmd,
stdout=sp.PIPE,
stderr=sp.PIPE,
shell=True,
bufsize=1,
universal_newlines=True,
) as p:
for line in p.stderr:
print(line, end="")
# convert nested samples to posterior samples
outpost = os.path.join(outdir, "{}_post.hdf".format(label))
runcmd = " ".join([n2p, "-p", outpost, outfile])
with sp.Popen(
runcmd,
stdout=sp.PIPE,
stderr=sp.PIPE,
shell=True,
bufsize=1,
universal_newlines=True,
) as p:
for line in p.stdout:
print(line, end="")
# get posterior samples
post = read_samples(outpost, tablename=LALInferenceHDF5PosteriorSamplesDatasetName)
lp = len(post["H0"])
postsamples = np.zeros((lp, len(priors)))
for i, p in enumerate(priors.keys()):
postsamples[:, i] = post[p.upper()]
# get evidence
hdf = h5py.File(outpost, "r")
a = hdf["lalinference"]["lalinference_nest"]
evsig = a.attrs["log_evidence"]
evnoise = a.attrs["log_noise_evidence"]
hdf.close()
# run bilby via the pe interface
runner = pe(
data_file=hetfile,
par_file=parfile,
prior=priors,
detector=detector,
outdir=outdir,
label=label,
)
result = runner.result
# evaluate the likelihood on a grid
gridpoints = 30
grid_size = dict()
for p in priors.keys():
grid_size[p] = np.linspace(
np.min(result.posterior[p]), np.max(result.posterior[p]), gridpoints
)
grunner = pe(
data_file=hetfile,
par_file=parfile,
prior=priors,
detector=detector,
outdir=outdir,
label=label,
grid=True,
grid_kwargs={"grid_size": grid_size},
)
grid = grunner.grid
# output comparisons
comparisons(label, outdir, grid, priors, cred=0.9)
# create results plot
allresults = {
"lalpulsar_parameter_estimation_nested": outpost,
"cwinpy_pe": result,
"cwinpy_pe (grid)": grid,
}
colors = {
key: plt.rcParams["axes.prop_cycle"].by_key()["color"][i]
for i, key in enumerate(allresults.keys())
}
plot = Plot(
results=allresults,
parameters=list(priors.keys()),
plottype="corner",
pulsar=parfile,
)
plot.plot(
bins=50,
smooth=0.9,
quantiles=[0.16, 0.84],
levels=(1 - np.exp(-0.5), 1 - np.exp(-2), 1 - np.exp(-9 / 2.0)),
fill_contours=True,
colors=colors,
)
plot.savefig(os.path.join(outdir, "{}_corner.png".format(label)), dpi=150)